Software support for SBGN maps: SBGN-ML and LibSBGN (Martijn P. van Iersel, Alice C. Villéger, Tobias Czauderna, Sarah E. Boyd, Frank T. Bergmann, Augustin Luna, Emek Demir, Anatoly Sorokin, Ugur Dogrusoz, Yukiko Matsuoka, Akira Funahashi, Mirit I. Aladjem, Huaiyu Mi, Stuart L. Moodie, Hiroaki Kitano, Nicolas Le Novère, and Falk Schreiber
Software support for SBGN maps: SBGN-ML and LibSBGN Bioinformatics 2012 28: 2016-2021. )
Warning: Unless you really like mapping and markup languages this is likely to be a boring story. If you do (and I do), it is the sort of thing you will print out and enjoy reading. Just so you know.
Abstract:
Motivation: LibSBGN is a software library for reading, writing and manipulating Systems Biology Graphical Notation (SBGN) maps stored using the recently developed SBGN-ML file format. The library (available in C++ and Java) makes it easy for developers to add SBGN support to their tools, whereas the file format facilitates the exchange of maps between compatible software applications. The library also supports validation of maps, which simplifies the task of ensuring compliance with the detailed SBGN specifications. With this effort we hope to increase the adoption of SBGN in bioinformatics tools, ultimately enabling more researchers to visualize biological knowledge in a precise and unambiguous manner.
Availability and implementation: Milestone 2 was released in December 2011. Source code, example files and binaries are freely available under the terms of either the LGPL v2.1+ or Apache v2.0 open source licenses from http://libsbgn.sourceforge.net.
Contact: sbgn-libsbgn@lists.sourceforge.net
I included the hyperlinks to standards and software for the introduction but not the article references. Those are of interest too but for the moment I only want to entice you to read the article in full. There is a lot of graph work going on in bioinformatics and we would all do well to be more aware of it.
The Systems Biology Graphical Notation (SBGN, Le Novère et al., 2009) facilitates the representation and exchange of complex biological knowledge in a concise and unambiguous manner: as standardized pathway maps. It has been developed and supported by a vibrant community of biologists, biochemists, software developers, bioinformaticians and pathway databases experts.
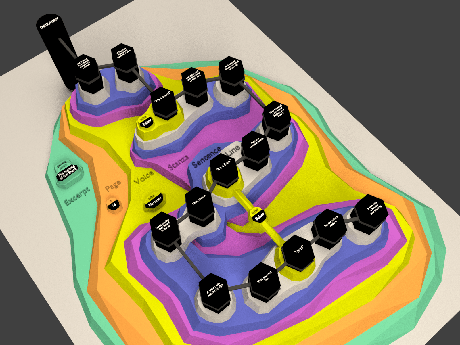
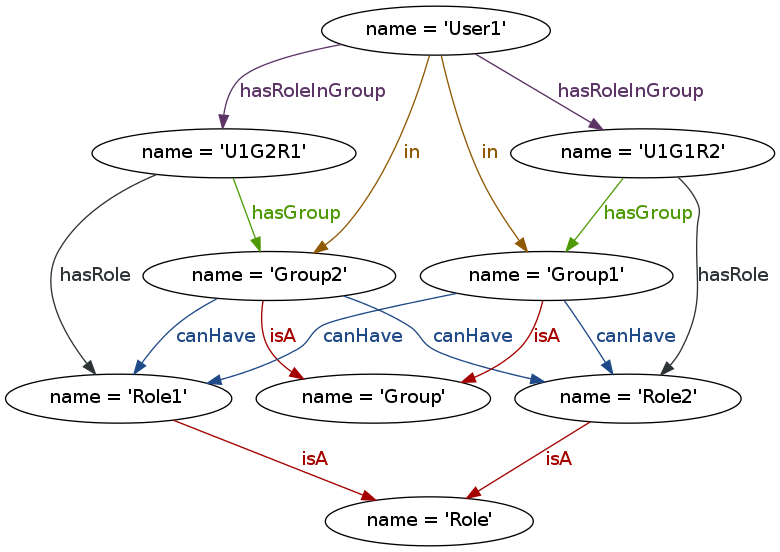
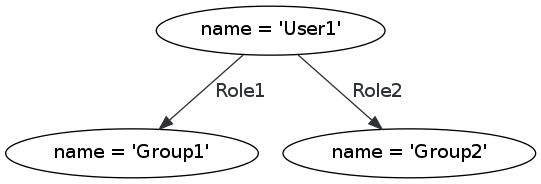
SBGN is described in detail in the online specifications (see http://sbgn.org/Documents/Specifications). Here we summarize its concepts only briefly. SBGN defines three orthogonal visual languages: Process Description (PD), Entity Relationship (ER) and Activity Flow (AF). SBGN maps must follow the visual vocabulary, syntax and layout rules of one of these languages. The choice of language depends on the type of pathway or process being depicted and the amount of available information. The PD language, which originates from Kitano’s Process Diagrams (Kitano et al., 2005) and the related CellDesigner tool (Funahashi et al., 2008), is equivalent to a bipartite graph (with a few exceptions) with one type of nodes representing pools of biological entities, and a second type of nodes representing biological processes such as biochemical reactions, transport, binding and degradation. Arcs represent consumption, production or control, and can only connect nodes of differing types. The PD language is very suitable for metabolic pathways, but struggles to concisely depict the combinatorial complexity of certain proteins with many phosphorylation states. The ER language, on the other hand, is inspired by Kohn’s Molecular Interaction Maps (Kohn et al., 2006), and describes relations between biomolecules. In ER, two entities can be linked with an interaction arc. The outcome of an interaction (for example, a protein complex), is considered an entity in itself, represented by a black dot, which can engage in further interactions. Thus ER represents dependencies between interactions, or putting it differently, it can represent which interaction is necessary for another one to take place. Interactions are possible between two or more entities, which make ER maps roughly equivalent to a hypergraph in which an arc can connect more than two nodes. ER is more concise than PD when it comes to representing protein modifications and protein interactions, although it is less capable when it comes to presenting biochemical reactions. Finally, the third language in the SBGN family is AF, which represents the activities of biomolecules at a higher conceptual level. AF is suitable to represent the flow of causality between biomolecules even when detailed knowledge on biological processes is missing.
Efficient integration of the SBGN standard into the research cycle requires adoption by visualization and modeling software. Encouragingly, a growing number of pathway tools (see http://sbgn.org/SBGN_Software) offer some form of SBGN compatibility. However, current software implementations of SBGN are often incomplete and sometimes incorrect. This is not surprising: as SBGN covers a broad spectrum of biological phenomena, complete and accurate implementation of the full SBGN specifications represents a complex, error-prone and time-consuming task for individual tool developers. This development step could be simplified, and redundant implementation efforts avoided, by accurately translating the full SBGN specifications into a single software library, available freely for any tool developer to reuse in their own project. Moreover, the maps produced by any given tool usually cannot be reused in another tool, because SBGN only defines how biological information should be visualized, but not how the maps should be stored electronically. Related community standards for exchanging pathway knowledge, namely BioPAX (Demir et al., 2010) and SBML (Hucka et al., 2003), have proved insufficient for this role (more on this topic in Section 4). Therefore, we observed a second need, for a dedicated, standardized SBGN file format.
Following these observations, we started a community effort with two goals: to encourage the adoption of SBGN by facilitating its implementation in pathway tools, and to increase interoperability between SBGN-compatible software. This has resulted in a file format called SBGN-ML and a software library called LibSBGN. Each of these two components will be explained separately in the next sections.
Of course, there is always the data prior to this markup and the data that comes afterwards, so you could say I see a role for topic maps.