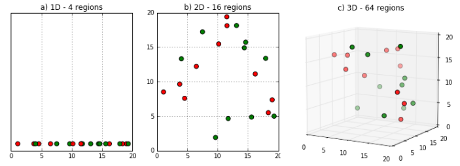
Wikipedia introduces its article on the Curse of Dimensionality with:
In numerical analysis the curse of dimensionality refers to various phenomena that arise when analyzing and organizing high-dimensional spaces (often with hundreds or thousands of dimensions) that do not occur in low-dimensional settings such as the physical space commonly modeled with just three dimensions.
There are multiple phenomena referred to by this name in domains such as sampling, combinatorics, machine learning and data mining. The common theme of these problems is that when the dimensionality increases, the volume of the space increases so fast that the available data becomes sparse. This sparsity is problematic for any method that requires statistical significance. In order to obtain a statistically sound and reliable result, the amount of data needed to support the result often grows exponentially with the dimensionality. Also organizing and searching data often relies on detecting areas where objects form groups with similar properties; in high dimensional data however all objects appear to be sparse and dissimilar in many ways which prevents common data organization strategies from being efficient.
The term curse of dimensionality was coined by Richard E. Bellman when considering problems in dynamic optimization.[1][2]
The “curse of dimensionality” is often used as a blanket excuse for not dealing with high-dimensional data. However, the effects are not yet completely understood by the scientific community, and there is ongoing research. On one hand, the notion of intrinsic dimension refers to the fact that any low-dimensional data space can trivially be turned into a higher dimensional space by adding redundant (e.g. duplicate) or randomized dimensions, and in turn many high-dimensional data sets can be reduced to lower dimensional data without significant information loss. This is also reflected by the effectiveness of dimension reduction methods such as principal component analysis in many situations. For distance functions and nearest neighbor search, recent research also showed that data sets that exhibit the curse of dimensionality properties can still be processed unless there are too many irrelevant dimensions, while relevant dimensions can make some problems such as cluster analysis actually easier.[3][4] Secondly, methods such as Markov chain Monte Carlo or shared nearest neighbor methods[3] often work very well on data that were considered intractable by other methods due to high dimensionality.
But dimensionality isn’t limited to numerical analysis. Nor is its reduction.
Think about the number of dimensions along which you have information about your significant other, friends or co-authors. Or any other subject, abstract or concrete, that you care to name.
However many dimensions you can name for any given subject, in human discourse we don’t refer to that dimensionality as the “curse of dimensionality.”
In fact, we don’t notice the dimensionality at all. Why?
We reduce all those dimensions into a name for the subject and that name is what we use in human discourse.
Dimensional reduction to names goes a long way to explaining why we get confused by names.
Another speaker has reduced a different set of dimensions (which are not shown as it were) to the same name that we use as the reduction of a different set of dimensions.
Sometimes the same name will expand into a different set of dimensions and sometimes different names expand into the same set of dimensions.
One of those dimensions being the context of usage, which when our expansion of the name doesn’t fit, prompts us to ask the speaker for one or more additional dimensions to identify the subject of discussion.
We do that effortlessly, reducing and expanding dimensions to and from names in the course of a conversation. Or when reading or writing.
The number of dimensions for any name increases as we know more about any given subject. Not to mention being impacted by our interaction with others who use the same name as we adjust, repair or change the dimensions we expand or reduce for any particular name.
Dimensionality isn’t a curse. The difficulties we associate with dimensionality and numeric analysis are a consequence of using an underpowered tool, that’s all.